
What's new in 3decision® 2021.1?
The updated 3decision® version has some important new features for cross-team collaboration and structure-based drug design project management. A lot of improvements are made on the structure visualization UI as well.

Spotting differences in large structure collections
This short post shows how to quickly analyze larger collections of structures and check for common and different parts among such large sets of structures.
What are the challenges for medicinal chemists when working with 3D structures? - Interview
The continuous increase in the available 3D structural data brings along changes in the medicinal chemistry field. Dr. George Sheppard, Senior Principal Research Scientist at AbbVie, shared his thoughts on the benefits for medicinal chemists using 3D information, current challenges and limitations, and proposed potential ways for improvement, in the following interview.


How to speed up the preparation of data sets for Structure-Based Drug Design? – Webinar Recap
The construction of accurate data sets prior to the in-silico experiments is a crucial first step in many SBDD projects. In our latest webinar, we explained how 3decision® can help ease and speed up this process. For better understanding, we presented three common use cases in SBDD using our platform.

Getting ready for AlphaFold with 3decision®
Google-owned company DeepMind has achieved to predict 3D protein shapes with high accuracy and speed using the AI program AlphaFold. Although still in the early phase, this method can have a significant impact on drug discovery, accelerating the development of new drugs. In this article, we are discussing the importance of getting ready for the new source of structural data and how to gain the most of structural knowledge.

Should medicinal chemists use 3D molecular modeling tools?
We sat down with Dr. Alexis Denis, Ph.D. Medicinal Chemistry Director at Galapagos to collect his thoughts on the question: Should medicinal chemists use 3D molecular modeling tools?
We were able to understand how he manages the collaboration in SBDD teams, but also to understand his vision of how these positions/roles could evolve in the future.
Discngine & UserStudio: Uplifting User Experience in life science research applications
Recently, our collaborative platform, 3decision, just saw itself complemented with two new features (subpocket search & interaction search) and a radically different User Experience (UX). This new UX, with an updated User Interface (UI), is the result of months of collaborative work between UserStudio and the 3decision team.
In order to tell the story of this project, we gathered people from both companies to answer a few questions.


3decision® v2 – Discover the interaction search
The highly requested and long-awaited interaction search is finally here! Given the great amount of structural data we have at hand, the ability to mine ligand-protein complexes based on the interactions they create was a missing piece in our analytics toolbox.

3decision® v2 – Discover the subpocket search
Select a couple of amino acids in the binding site of your protein and retrieve, in a matter of minutes, other structures in the database which contain a similar subpocket. Similar meaning, similar features arranged similarly in 3D space. This is what the new subpocket search feature will allow you to do in the next release of 3decision!

3decision® v2 – Discover the new UI
Last year, we set out a process to improve the user experience in 3decision. After years of gathering feedback and interviewing key users, we knew it was time to address the design and workflow flaws in the user interface. With the help of User Studio, an external UI/UX design company, we went through a deep refactoring and restyling of the front-end. The resulting work will be available in the next upcoming release of 3decision (3decision v2)!

Discngine announces that Sosei Heptares will use its 3decision® software to create an unprecedented structural GPCR chemogenomics platform
Discngine announces that Sosei Heptares will use its 3decision® software to create an unprecedented structural GPCR chemogenomics platform. 3decision® will enable Sosei Heptares to combine public and proprietary structural data and derive unique GPCR structural insights.

SARS-Cov-2 - part 3 - nsp1: A hopefully more detailed analysis of the cellular saboteur
My German cartesian mind (here we go with stereotypes) asks me to start with the first protein expressed by the viral genome of SARS-Cov-2, nsp1. That is, by far, not the low hanging fruit. But it is interesting to learn more about the roles of the different constituents of the virus. Nsp1 actually has also a few very interesting roles to facilitate the viral lifecycle in its host environment.
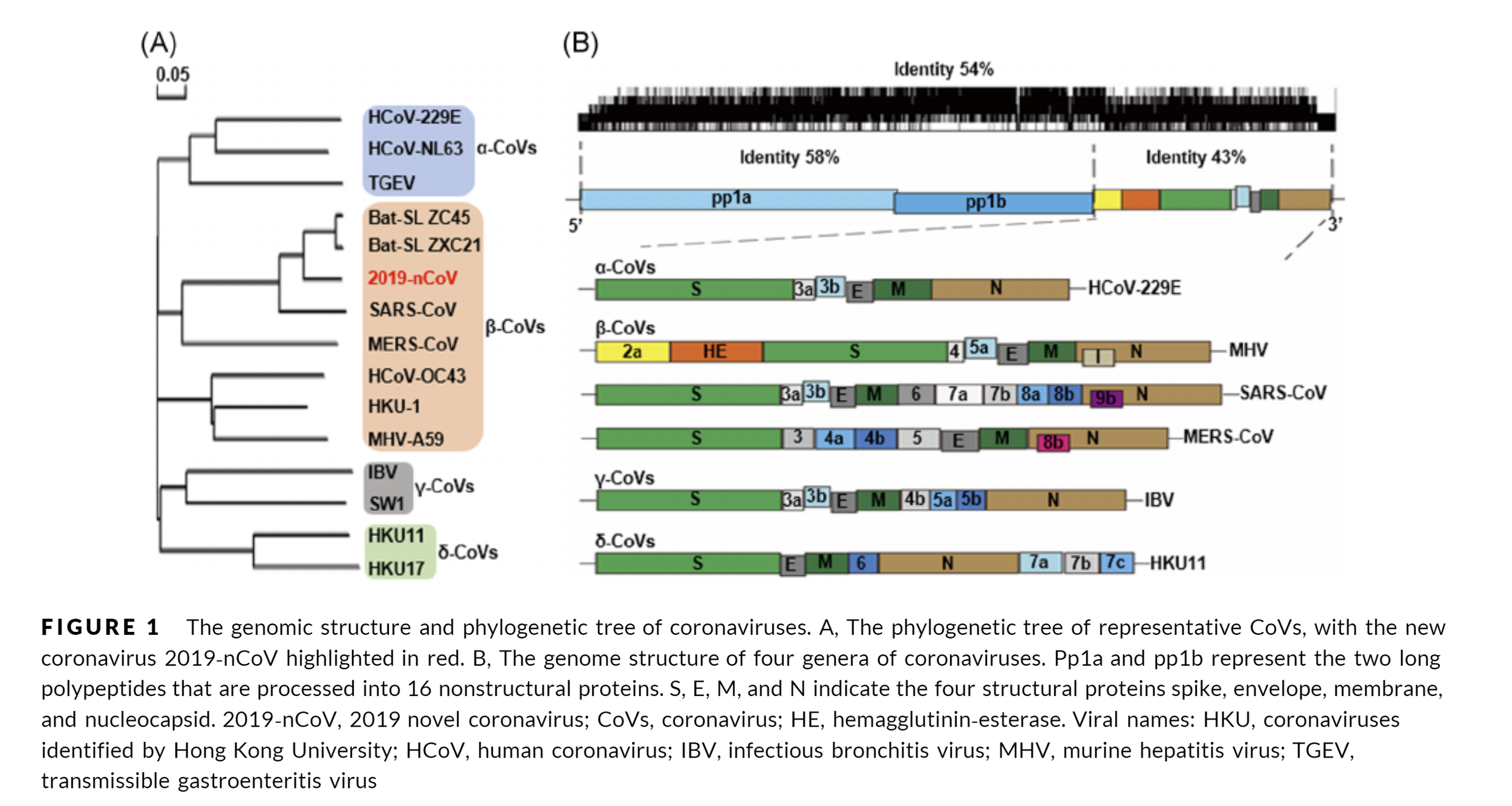
SARS-CoV-2 - part 2 - From the viral genome to protein structures
The bread and butter of every structure based drug design or drug ID effort is the need for 3D structural information of a target of interest related to the phenotypic outcome we want to alter. In the case of SARS-Cov-2 several experimental structures are already available in the RCSB PDB. Other a large scale fragment screens from the Diamond Light Source are slowly becoming available (available in the 3decision protease project for now) on public resources likes the RCSB PDB. Popular homology modelling services also already started to provide several high quality models for some of the yet not resolved structures of the viral proteome.
But let’s first have a look what the viral genome actually contains and what are currently pursued efforts.

SARS-CoV-2 - part 1 - Thriving for a systematic target and hit ID effort
Gabriella outlined in previous posts (part I & part II) a few things about SARS-Cov-2. Back at the time the crisis was still far away from our daily lives but things have changed dramatically during the last weeks. After our initial communication to make 3decision.discngine.cloud freely available for all Covid-19 related collaboration projects we were contacted independently by several volunteers from academic and industrial groups to collaborate on a global effort. At the mean time a lot of novel initiatives have seen the day and everybody is publishing blog posts (like I am right now), linkedIn articles, chemarxiv or bioarxiv preprints etc. It’s a little bit a scientific wild-west happening right now on a global scale.
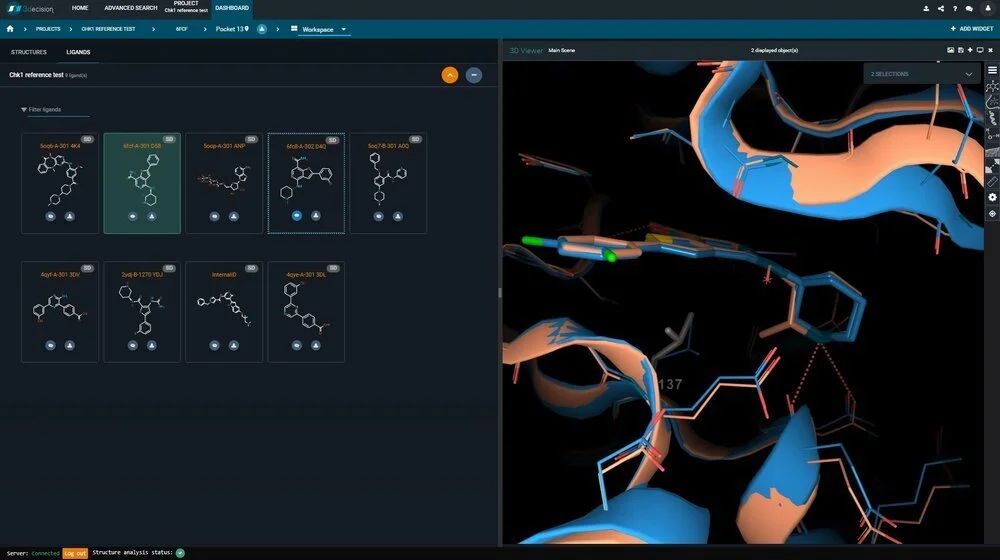
Discover 3decision® - Part 2: Creating a collaborative platform for Structure-Based Drug Discovery
As we explained in our previous article, the number of biomolecular structures available for computational analysis is increasing daily, which, for SBDD translates to ever-greater opportunities for discovery.
But wait, is that actually true?
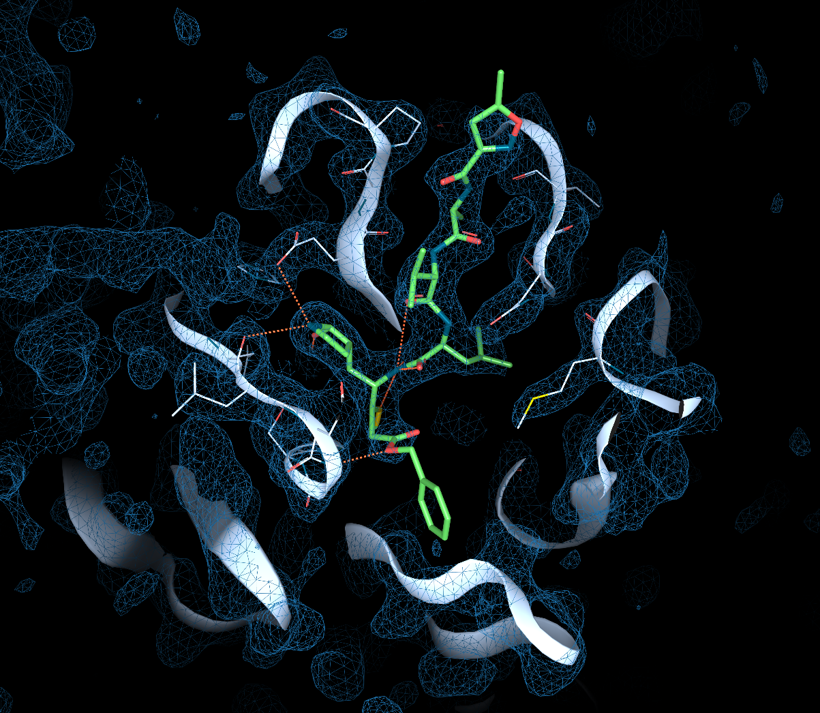
2019-nCov, structurally speaking - part II
As a follow-up to my previous article 2019-nCoV, structurally speaking, I thought I’d share some of the news on the protein structure side and some insights from the structural analyses we’ve done at Discngine.

2019-nCov, structurally speaking
I guess nobody has missed the fact that we are in the middle of a new Coronavirus outbreak. In the beginning, I followed the updates from far, without going in too much detail. That was before I saw a LinkedIn post on a homology model of a 2019-nCoV protease last week.
